Evolutionary Dynamics
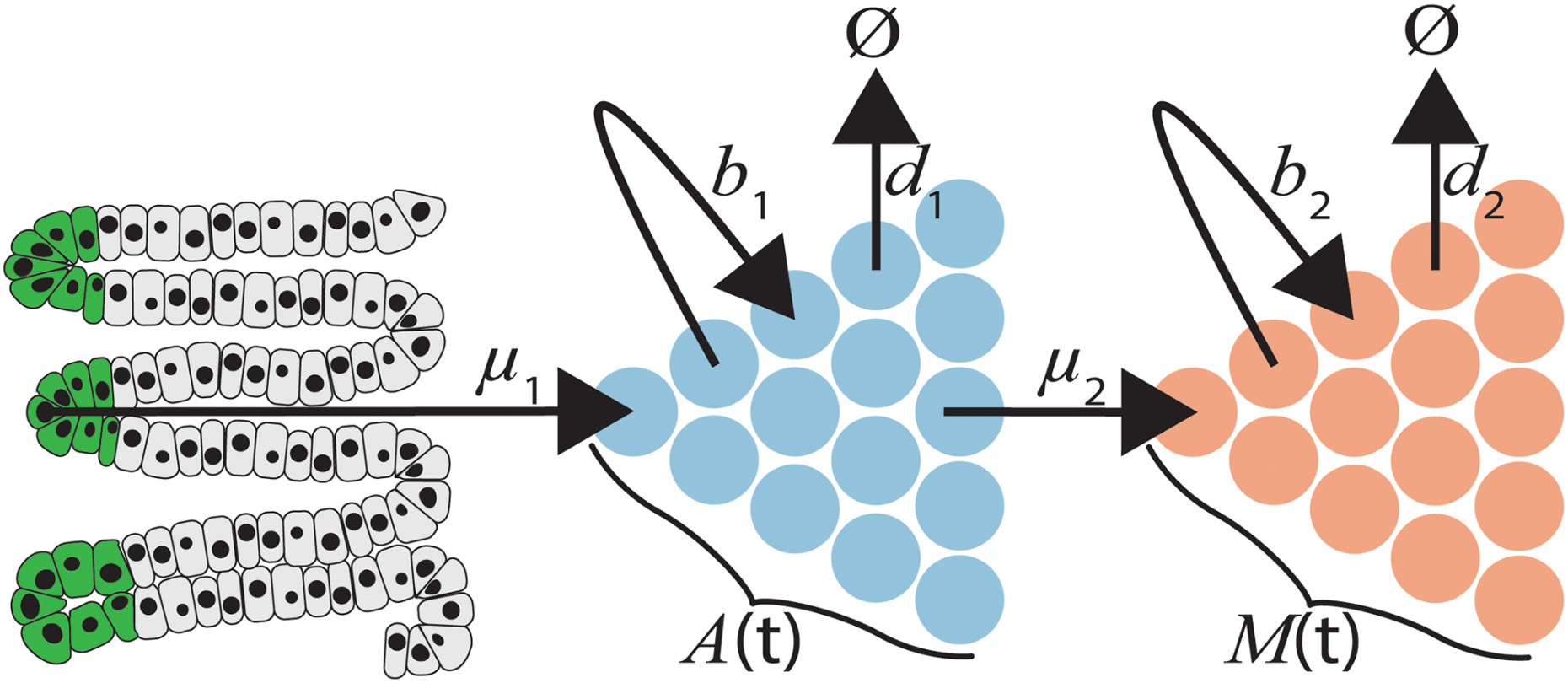
Evolutionary change is a hallmark of biological systems. We are interested in the evolutionary dynamics of pathogen populations, specifically RNA viruses, such as HIV and HCV, and tumor cell populations. In these systems, evolutionary changes occur on a time scale that is observable and the evolutionary dynamics are responsible for clinical outcomes of the disease. Using evolutionary modeling and population genetics, we have modeled, for example, the intra-host evolution of HIV-1 and the development of colorectal cancer.
Many disease-causing evolutionary processes of pathogenic agents occur repeatedly, for example, viral infections of their hosts. Data from such reproducible evolutionary processes can be used to learn structural constraints on the available evolutionary pathways and on the dynamics of genetic adaptation. We develop probabilistic graphical models to describe the accumulation of genetic alterations. Specifically, we predict the development of drug resistance in HIV and the occurrence of cancer-driving mutations in tumors.
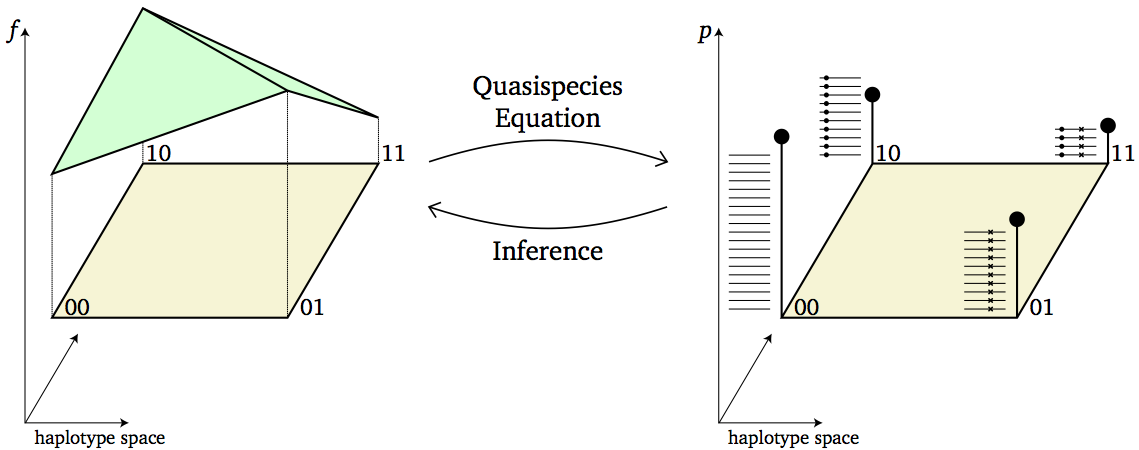
A major goal of our research is to use DNA sequencing data obtained from evolving populations to infer models and parameters of the underlying evolutionary process. For example, employing the quasispecies model of viral evolution, we have constructed a computational method for inferring the fitness of the individual strains in the population. For cancer, we have developed computational methods for reconstructing the evolutionary history of a tumor from single-cell sequencing data.
Selected references:
- external pageInferring genetic interactions from comparative fitness datacall_made
Crona et al., eLIFE, 2017 - external pagepathTiMEx: Joint Inference of Mutually Exclusive Cancer Pathways and Their Progression Dynamicscall_made
Cristea et al., J Comput Biol, 2017 - Predicting colorectal cancer risk from adenoma detection via a two-type branching process model
Lang et al., PLoS Comput Biol, 2020